
Thomas Squire
Download a printable version of this document here.
A proteins three dimensional structure is complex, however it can be broken down to allow a greater understanding of how a protein folds into its complex three dimensional structure.
Primary (1°) structure
The primary protein structure is the simplest way to describe a protein, and is simply the linear sequence of amino acids that makes up the peptide chain. Although the primary structure maybe simple, it is key in determining how the protein folds as it is the interactions between amino acids that govern subsequent folding.
Secondary (2°) Structure
The secondary protein structure describes the first step in how the amino acid sequence starts to fold in three dimensional space. There are two main secondary structures that occur, both of which are governed by hydrogen bonding between carbonyl and amino groups of the amino acids in the peptide chain. These are alpha helices and beta sheets. It should be noted that a single secondary structure rarely extends throughout the peptide chain, it usually just forms a section within the chain.

Alpha helices
The alpha-helix structure is in the form of a right handed coil and is formed through hydrogen bonding between the carbonyl group and amino group of every 3.6 amino acids. This along with the fact all natural amino acids are in the form of 1 enantiomer* leads to the right handed helical structure forming (* with the exception of glycine which is not chiral). Note the R groups of the amino acids point out from the α-helix and are not involved in the hydrogen bonds that maintain the structure.
Beta sheets
Beta sheets (sometimes described as beta pleated sheets) are formed when the linear chains fold back on themselves leaving the chains to run parallel to each other. This allows hydrogen bonds to form between carbonyl and amino groups of the parallel linear chains, therefore forming the beta sheets. Beta sheets are often denoted as parallel arrows (see image) identifying the direction the sequence runs, as beta sheets can both be parallel or anti parallel depending on how the N- and C-termini align (i.e. either running in same or opposite direction). Note the R groups of the amino acids in the beta sheet point out at a right angle to the hydrogen bonds holding the beta sheets together and not involved in the hydrogen bonds that maintain the structure.
Tertiary (3°) Structure
The tertiary protein structure describes how the secondary structure elements interact together to form a complex three dimensional protein structure (also known as a domain). These interactions are generally between the R groups of the amino acids and include many non covalent interactions; hydrogen bonding, ionic bonding (forming salt bridges), dipole-dipole interactions, and London dispersion forces. Disulphide bridges can also form through a covalent bond between two sulphur atoms of the cysteine R group.
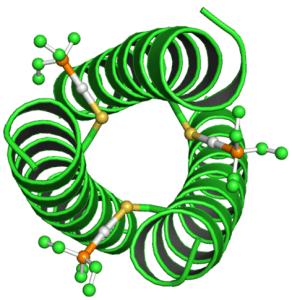
The manner in which hydrophilic / hydrophobic portions of the peptide chain interact is also highly important in determining the tertiary protein structure and is often the main driving force behind folding. For example the majority of proteins are present in an aqueous environment, thus it is important for the tertiary structure to fold allowing the hydrophilic R groups to interact with the aqueous environment while shielding the hydrophobic R groups within the protein. An example of this can be seen in how a coiled coil forms. The secondary alpha helices structures come together to allow a hydrophobic pocket to form containing hydrophobic R groups of the alpha helix, while also exposing the hydrophilic R groups to the aqueous environment surrounding the coiled coil.
Quaternary (4°) Structure
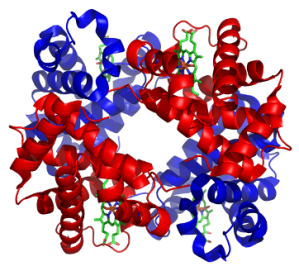
The quaternary structure of proteins describes how multiple protein subunits (or domains) can come together to form a larger protein complex (i.e. a protein with multiple domains). Proteins are often characterised as being one massive polypeptide, however this disregards the quaternary structures ability to bring together multiple domains. In fact many proteins are the product of multiple domains coming together such as the metalloprotein haemoglobin (right) with its four domains.
To learn more about protein structures, see the chemBAM experiment page – protein building blocks.

This work is licensed under a Creative Commons Attribution 4.0 International License.